Are computer simulations the future of rapid COVID-19 drug development?
Posted: 11 October 2021 | Navodya Roemer (University of Lincoln) | No comments yet
Researchers have visualised SARS-CoV-2 protein dynamics using in silico methods. In this article, Navodya Roemer explains how a team from the University of Warwick developed a computational strategy that could assist scientists in the production of new treatments and drugs for COVID-19.

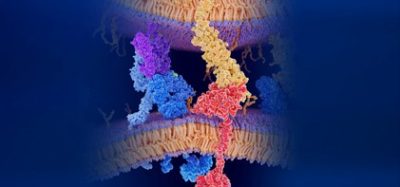
In recent years, drug discovery supported by computer modelling has become a viable method to corroborate experimental efforts in the large-scale screening of prospective therapeutic compounds. This approach is part of the overarching theme of bioinformatics which aims to harness the power of modern computational techniques for the in silico design of medicines. The advantage of this strategy lies not only in an increased screening speed, but also in its predictive power for novel drug development. In early 2020, protein structures related to SARS-CoV-2 began to appear in the international protein data bank (PDB). Such structures provided the precise location of the atoms in each protein, forming the basis for an understanding on how their biological mechanisms might function. Chief among these structures were the protease, important in the life cycle of SARS-CoV-2 and the Spike (S) glycoprotein, an integral protein protruding out of the viral capsid which the virus uses when attaching itself to a human cell during infection.
To understand the biological function of a given structure such as the S protein, it is important to know how such a structure moves. Experimentally, this is very hard to ascertain and requires months of careful experimentation. Computational modelling, on the other hand, can provide crucial information at an accelerated rate. However, in making such models, simplified assumptions must be made which can reduce the accuracy of the results that are obtained. In our work, we used an approach in which a protein structure is divided into rigid and flexible parts. This allowed us to rapidly model the motion of the protein by keeping the rigid parts stable while allowing for the movement of the flexible regions.1

Although our method as described above is rapid, employing it to study the more than 300 SARS-CoV-related protein structures that had become available in the PDB by mid-2020 required substantial computational effort. Even saving the resulting dynamics in their compressed formats still required hundreds of gigabytes. Furthermore, we also discovered the occasional problem in the deposited PDB structures where due to the complexity of proteins – the SARS-CoV-2 S protein contains 2,995 protein “residues”, equivalent to a certain number of atoms – sometimes a lack of spatial resolution prohibited the application of our approach. Overall, we were able to successfully analyse the dynamics of 297 SARS-CoV-2 related proteins.
Having identified the precise nature of the dynamics for the S protein, scientists around the word can now use their own in silico design methods to quickly screen for existing and novel drug molecules that might stop the dynamics – in effect, similar to tying down the RBD to prevent any binding to human cells.
Our work on the SARS-CoV-2 S protein concentrated on the original structures emerging in early 2020. Since then, mutations of the virus have emerged which are predominantly appearing on the S protein. This clearly highlights the importance of the S protein as a route towards infection but also as a potential target for vaccines and suppression of infective mechanisms.
…drug discovery supported by computer modelling has become a viable method to corroborate experimental efforts”
We are now using the knowledge gained from our study to understand how the mutations in the S protein might affect its dynamics. In March 2021, the first scientific results came out that provided PDB structures of the SARS-CoV-2 mutations for the UK (alpha), South African (beta) and Brazilian (gamma) variants. Structures for other variants, including the much-discussed delta variant, are not yet known and we have to construct models for these ourselves. First results from our modelling suggest that the dynamical properties do not change very much – this should provide some comfort as it might indicate that vaccines based on the shape of the S proteins will continue to work. Clearly, much more work is needed here to be able to precisely describe each mutant variation.
The full repository of all dynamics is publicly available at the University of Warwick Flex-Covid19 data repository, here.
About the author
Navodya Roemer is a 2021 graduate student from the University of Lincoln where she obtained a bachelor’s degree in biomedical sciences. Having experienced the Covid-19 pandemic both within the lecture hall and outside of it, she became interested in how she could support the scientific community in combating this pandemic. During lockdown at home, she and her dad, a physicist at Warwick University, teamed up with Warwick colleague Katrine Wallis from life sciences to model protein dynamics for SARS-CoV-2 structures as described above. Novo is hoping to continue her studies and is at the moment eagerly awaiting admission results from various German universities where she is aiming to pursue a master’s in life sciences.
References
- Römer, R, Römer, N, & Wallis, A (2021). Flexibility and mobility of SARS-CoV-2-related protein structures. Scientific Reports, 11, 4257. https://doi.org/10.1101/2020.07.12.199364.
Related topics
Big Data, Drug Development, Informatics, Molecular Targets, Protein, Proteomics, Vaccine
Related conditions
Covid-19
Related organisations
Warwick University